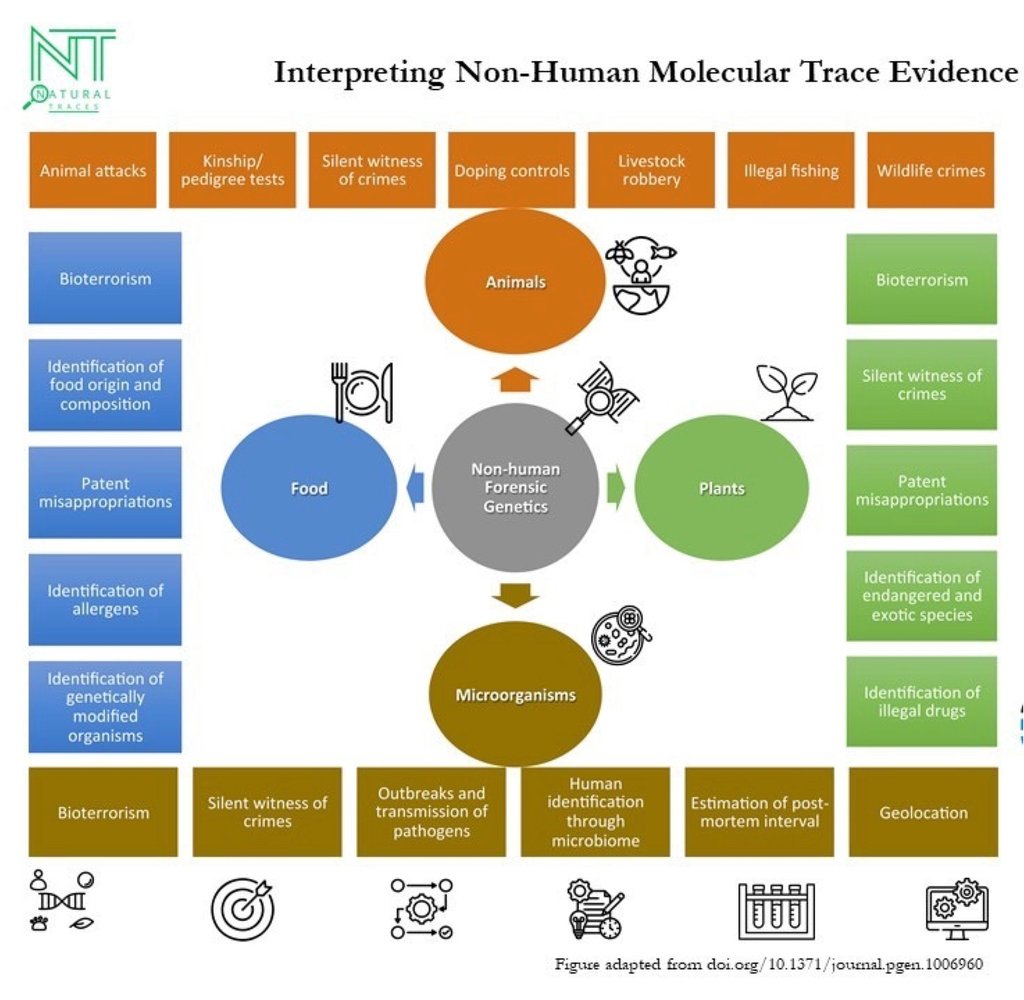
Interpreting Non-Human Molecular Trace Evidence
Non-Human Forensic Genetics
31 August – 3 September 2026
i3S - Instituto de Investigação e Inovação em Saúde, Institute for Research and Innovation in Health, Universidade do Porto, Portugal
Nádia Pinto
Programme
Nadia Pinto
Filipe Pereira, Pedro Ferreira, Eran Elhaik, Henk Braig
Arrival: Sunday, 30 August
Day 1, 31 August, 09h – 17h
Non-Human Forensic Genetics: Methodologies, Applications, Exceptions, and Case Studies
Nádia Pinto, i3S-Institute for Research and Innovation in Health, University of Porto, Portugal, and Henk Braig, National University of San Juan (UNSJ), San Juan, Argentina
Foundations and scope
Basic concepts in non-human forensic genetics
Why non-human genetic material matters in forensic contexts
When the field behaves like human forensic genetics and when it does not
Methodologies
Sampling, extraction, quantification, and validation
Marker types
Software and laboratory workflow, including sequence curation and quality control
Interpretation
Database-based identification and its limits
Statistical evaluation of evidence
Phylogenetic and evolutionary inference, including recombination and model choice
Applications
Zoology, botany, microbiology, and food forensics
Species identification, individualization, kinship, population assignment, outbreak tracing, and traceability
Exceptions and pitfalls
Nonstandard or difficult cases
Low-quality databases, poor population structure, degraded samples, mixtures, and validation limits
Situations where the usual forensic assumptions do not hold
Case studies
Day 2, 01 September, 09h – 17h
Molecular Identification of Non-Human Biological Evidence: A Comprehensive Workflow (hands-on) by
Filipe Pereira, Department of Genetics and Biotechnology (DGB), University of Trás-os-Montes and Alto Douro, Portugal
Basic concepts related to biological identifications using DNA
Overview of DNA sequencing in the laboratory
Introduction to software for DNA sequence analyses
Installation and use of the Geneious software for DNA sequence management
Sequence curation and quality control (electropherograms, formats)
Brief overview of alignment of DNA sequences and phylogenetic inferences
Database-based identifications (BLAST, BOLD): strengths and pitfalls
Examples of real cases of using DNA for biological identifications
Data interpretation and reporting
Day 3, 02 September, 09h – 17h
Introduction to Biological Sequence Databases Pedro G. Ferreira, Computer Sciences Department (hands-on), Faculty of Sciences of the University of Porto, Portugal
Sequence data formats and metadata.
Types of biological sequence databases (primary/non-curated and secondary/curated).
How sequences are organized, annotated, and linked.
Retrieving sequences efficiently.
Database quality assessment and common pitfalls.
Using sequence databases in non-human forensic genetics and metagenomics.
Introduction to Metagenomic Data Analysis: From Raw Sequencing Reads to Microbial Profiles (hands-on), Eran Elhaik, Ancient DNA Hub, Lund, Sweden
Introduction to Metagenomics and Microbiome Research
§ What metagenomics is and why it is used
§ Difference between shotgun sequencing and amplicon sequencing
§ Applications in ecology, health, and forensic science
Understanding Sequencing Data
What sequencing reads are
Structure of FASTQ files
Paired-end sequencing and quality scores
Quality Control and Data Cleaning
Why sequencing data must be cleaned
Identifying sequencing errors and contaminants
Read trimming and filtering
Example tools: FastQC, MultiQC, fastp, Trimmomatic
Day 4, 03 September, 09h – 17h
Introduction to Metagenomic Data Analysis: From Raw Sequencing Reads to Microbial Profiles – continuation, Eran Elhaik
Preparing Reads for Analysis
Handling paired-end reads
Removing host DNA contamination
Mapping reads to reference genomes
Example tools: repair.sh, Bowtie2, Samtools
Microbial Identification and Classification
Assigning sequencing reads to microbial taxa
How reference databases are used
Generating microbial abundance tables
Example tools: Kraken2 and Bracken
Amplicon Analysis and Microbial Community Profiles
Processing amplicon sequencing data
Merging reads and removing chimeras
Clustering sequences and generating OTU tables
Example tools: PANDAseq and VSEARCH
Workshop’s challenge
Workshop Dinner
Departure: Friday, 4 September




















Logistics


You must bring your own laptop!
Venue
The workshop will be held at i3S, organized with support from the i3S Advanced Training Unit.
i3S-Institute for Research and Innovation in Health, University of Porto
Rua Alfredo Allen, 208
4200-135 Porto, Portugal
GPS coordinates: 41º 10’ 30.008’’ N, 8º 36’ 12.488’’ W.


